Trends in daily R package downloads
August 23, 2016 Code R CRAN trend forecasting
This post was prompted by this blog about using the cranlogs package by Gabor Csardi. But my own interest as long time package developer dates back to this post by Ben Bolker. I like to see that my packages are being used. So I thought why stop at counting downloads and plotting the past. Why not predict into the future?
The R language has seen steady growth over the past decade in terms of its use relative to other programming languages and the number of available extension packages, both reflecting increasing community engagement. It is thus interesting to have a look at how individual packages perform over time based on CRAN download statistics.
As the R user community expands, the disparity between more and less popular packages is expected to increase. A handful of packages are on top of the CRAN download chart harvesting most of the attention of novice R users. However, popularity and trending based on downloads is not necessarily the best measure of overall impact, more like a measure that is easiest to quantify. At least I like to think that 100 downloads with 10 users is way better than 100 downloads with a single user. These two scenarios are hard to tell apart though. So for now, let’s assume that download statistics reflect real impact.
So I wanted to look at temporal patterns in the past 3 years’ download
statistics from RStudio CRAN mirror
for individual R packages. I quantified past trend based
on fitting a log-linear glm model to counts vs. dates.
Then I used the forecast package by Rob Hyndman to make short-term forecast
based on an additive non-seasonal exponential smoothing model. Here is the plot_pkg_trend function:
library(cranlogs)
library(forecast)
plot_pkg_trend <-
function(pkg)
{
op <- par(mar = c(3, 3, 1, 1) + 0.1, las = 1)
on.exit(par(op))
## grab the data
x <- cran_downloads(pkg, from = "2013-08-21", to = "2016-08-19")
x$date <- as.Date(x$date)
x$std_days <- 1:nrow(x) / 365
## past trend
m <- glm(count ~ std_days, x, family = poisson)
tr_past <- round(100 * (exp(coef(m)[2L]) - 1), 2)
## future trend
s <- ts(x$count)
z <- ets(s, model = "AAN")
f <- forecast(z, h=365)
f$date <- seq(as.Date("2016-08-20"), as.Date("2017-08-19"), 1)
tr_future <- round(100 * (f$mean[length(f$mean)] / f$mean[1L] - 1), 2)
## plot
plot(count ~ date, x, type = "l", col = "darkgrey",
ylab = "", xlab = "",
ylim = c(0, quantile(x$count, 0.999)),
xlim = c(x$date[1L], as.Date("2017-08-19")))
lines(lowess(x$date, x$count), col = 2, lwd = 2)
polygon(c(f$date, rev(f$date)),
c(f$upper[,2L], rev(f$lower[,2L])),
border = NA, col = "lightblue")
lines(f$date, f$mean, col = 4, lwd = 2)
legend("topleft", title = paste("Package:", pkg), bty = "n",
col = c(2, 4), lwd = 2, cex = 1,
legend = c(paste0("past: ", tr_past, "%"),
paste0("future: ", tr_future, "%")))
## return the data
invisible(x)
}
Next, we list the top 10 R packages from the last month:
cran_top_downloads("last-month")
## rank package count from to
## 1 1 Rcpp 221981 2016-07-23 2016-08-21
## 2 2 digest 181333 2016-07-23 2016-08-21
## 3 3 ggplot2 170577 2016-07-23 2016-08-21
## 4 4 plyr 163590 2016-07-23 2016-08-21
## 5 5 stringi 154918 2016-07-23 2016-08-21
## 6 6 stringr 153917 2016-07-23 2016-08-21
## 7 7 jsonlite 152040 2016-07-23 2016-08-21
## 8 8 magrittr 139596 2016-07-23 2016-08-21
## 9 9 curl 134026 2016-07-23 2016-08-21
## 10 10 scales 130887 2016-07-23 2016-08-21
The top package was Rcpp by Dirk Eddelbuettel et al. and here is the daily trend plot with percent annual trend estimates:
plot_pkg_trend("Rcpp")
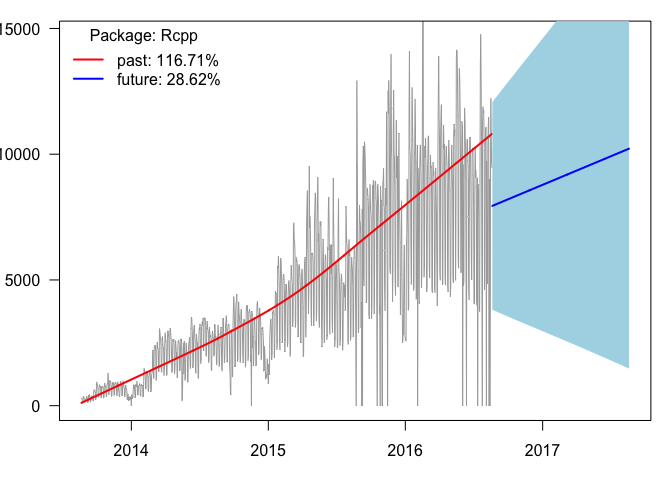
The increase is clear with increasing day-to-day variation
suggesting that the log-linear model might be appropriate.
Steady increase is predicted (the blue line starts at the
last observed data point, whereas the
red line is a locally weighted lowess smoothing curve on
past data).
The light blue 95% prediction
intervals nicely follow the bulk of the zig-zagging in the
observed trend.
Now for my packages, here are the ones I use most often:
op <- par(mfrow = c(2, 2))
plot_pkg_trend("pbapply")
plot_pkg_trend("ResourceSelection")
plot_pkg_trend("dclone")
plot_pkg_trend("mefa4")
par(op)
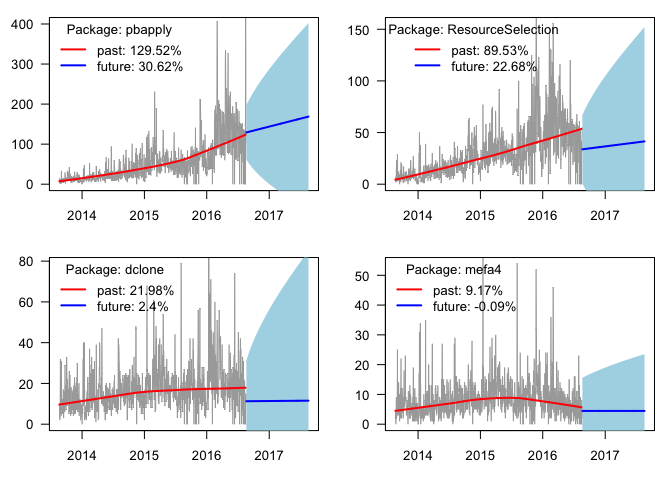
Two of them increasing nicely, the other two are showing some leveling-off. And finally, results for some packages that I am contributing to:
op <- par(mfrow = c(2, 2))
x <- plot_pkg_trend("vegan")
x <- plot_pkg_trend("epiR")
x <- plot_pkg_trend("adegenet")
x <- plot_pkg_trend("plotrix")
par(op)
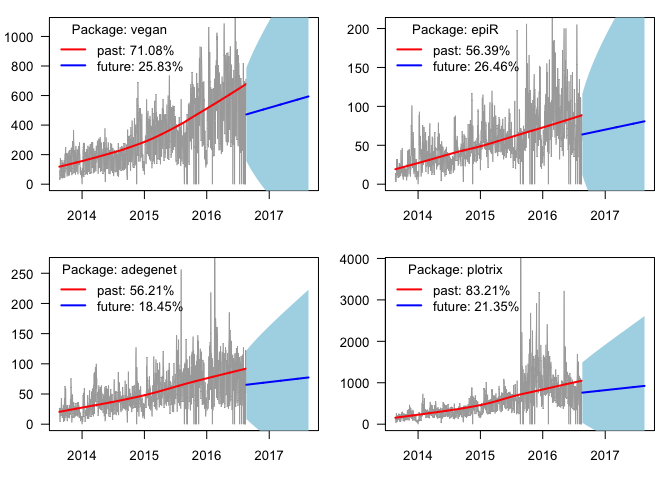
My approach as outlined here works for packages that have been with us since 2013. However, there are packages that went extinct (orphaned, excluded from CRAN, etc.) or appeared after 2013. These need to be handled with care when comparing across many packages or changing the time horizon. If you want to take the idea presented here a step further, you can compare observed and predicted increase in downloads for many packages, or compare recent download stats and trends. Let me know what you think in the comments below!
Closing the gap between data and decision making
Analyzing Ecological Data with Detection Error
This course is for researchers who analyze field observations. These data are often inconsistent because sampling protocols can change across projects, over time, or when older data are combined with new recordings.
- Analyzing Ecological Data with Detection Error
- Fitting removal models with the detect R package
- Shiny slider examples with the intrval R package
- Phylogeny and species traits predict bird detectability
- PVA: Publication Viability Analysis, round 3
ABMI (7) ARU (1) Alberta (1) BAM (1) C (1) CRAN (1) Hungary (2) JOSM (2) MCMC (2) PVA (2) PVAClone (1) QPAD (3) R (21) R packages (1) abundance (1) bioacoustics (1) biodiversity (1) birds (2) course (2) data (1) data cloning (4) datacloning (1) dclone (3) density (1) dependencies (1) detect (3) detectability (4) footprint (3) forecasting (1) functions (3) intrval (4) lhreg (1) mefa4 (1) monitoring (2) pbapply (5) phylogeny (1) plyr (1) poster (2) processing time (2) progress bar (4) publications (2) report (1) sector effects (1) shiny (1) single visit (1) site (1) slider (1) slides (2) special (3) species (1) trend (1) tutorials (2) video (4) workshop (2)